Study of enzymatic mechanisms
The increasing computing capacity of today’s computers and our deeper understanding of enzyme structures allow us to calculate enzymatic mechanisms by molecular dynamic simulations and quantum chemical methods. Such approaches enable to work on systems experimentally hard to tackle. Calmodulin, chymotrypsin, trypsin etc. were enzyme the inhibitors of which was studied by NMR-spectroscopy and X-ray diffraction in our group, complemented by MD and QM a decade ago.
More recently, Pseudouridine synthase responsible for the uridine–pseudouridine formation was modelled as well. Pseudouridylation affects several types of RNAs, linked to severe diseases (e.g. cancer, dyskeratosis congenital).
The mutation dependent pathogenicity of podocin can be related to the autosomal-recessive disorder of steroid-resistant nephrotic syndrome type 2. Thus, conformation and dimerization studies of various podocin mutants were completed early on by MD.
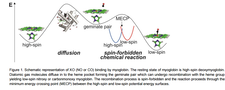
Ligand binding (CO, NO and O2) of hemeproteins (e.g. myloglobin) via MD simulations were performed to address the question of diffusion of ligands through the enzyme.
Kapcsolódó publikációk
-
Kálmán Tory , Dóra K. Menyhárd , Stéphanie Woerner , Fabien Nevo , Olivier Gribouval , Andrea Kerti , Pál Stráner , Christelle Arrondel , Evelyne Huynh Cong , Tivadar Tulassay , Géraldine Mollet , András Perczel , Corinne Antignac
Mutation-dependent recessive inheritance of NPHS2-associated steroid-resistant nephrotic syndrome
Nat. Genet. 46:299–304 | DOI: 10.1038/ng.2898 | PMID: 24509478 (2014) Kivonat -
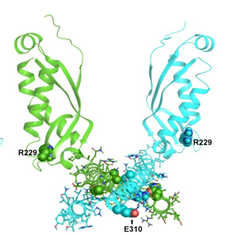
Ágnes Mikó , Dóra K. Menyhárd , Ambrus Kaposi , Corinne Antignac , Kálmán Tory
The mutation-dependent pathogenicity of NPHS2 p.R229Q: A guide for clinical assessment
Wiley Online Library 1854-1860 | DOI: 10.1002/humu.23660 (2018) Kivonat -
Dóra J. Kiss , Julianna Oláh , Gergely Tóth , Dóra K. Menyhárd , György G Ferenczy
Quantum chemical calculations support pseudouridine synthase reaction through a glycal intermediate and provide details of the mechanism
Springer Berlin Heidelberg 137-162 | doi.org/10.1007/s00214-018-2361-6 (2018) Kivonat -
Anikó Lábas , Dóra K. Menyhárd , Jeremy N. Harvey , Julianna Oláh
First principles calculation of the reaction rates for ligand binding to myoglobin: the cases of NO and CO
Wiley Online Library 5350-5358 | doi.org/10.1002/chem.201704867 (2017) Kivonat