Proteases of the complement system
The complement system plays a crucial role in the innate immunity, connected to the adaptive immune response too. Its action is based on regulated protease cascades, where proteases cleave and activate each other’s proenzyme forms in a strictly ordered manner. In cooperation with the Laboratory of Structural Biophysics of RCNS HAS and Directed Evolution Research Group of the Department of Biochemistry ELTE, our aim is to explore the structural bases of the function of various complement proteins.
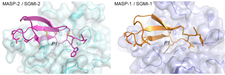
We studied structures of the early proteases of the classical and lectin pathways of complement activation using crystallographic methods in order to characterize their specificity and interactions at atomic level. The first enzymatic event of their cascades is the autoactivation of the first enzymes of the lectin and classical pathways – making these enzymes special because most serine proteases in the family are inactive in their proenzyme form. Based on the structure of C1r, the first enzyme of the classical pathway, we revisited the mechanism of C1r autoactivation. We suggest a “split-and-reassembly” mechanism, where a minor mechanic effect is enough to destabilize the head-to-tail dimer and initiate autoactivation. C1-inhibitor is a major downregulator of inflammatory processes in blood, being the inhibitor of also C1r, C1s, MASP-1 and MASP-2. Based upon the structure of its serpin domain we suggest a novel ”sandwich mechanism” explaining the effect of polyanions, such as heparin on the inhibitory affinity of C1-inhibitor. Comparison of the structures activated (cleaved) and proenzyme (single chain) forms of the catalytic regions of MASP-1 and MASP-2, the first enzymes of the lectin pathway helped to understand the structural conditions of autoactivation. Understanding the substrate specificities of these two enzymes was one our aim. Using highly specific inhibitors evolved using the phage-display technique, we solved the first enzyme-substrate complex-like structures of both MASP-1 and MASP-2. The 3D-structures explored structural elements of sub-site specificities, including an allosteric effect (PDB codes: 2oay, 2qy0, 4igd, 3gov, 1zjk, 1qx3, 3tvj, 4djz).
Related publications
-
József Dobó , Veronika Harmat , László Beinrohr , Edina Sebestyén , Péter Závodszky , Péter Gál
MASP-1, a Promiscuous Complement Protease: Structure of Its Catalytic Region Reveals the Basis of Its Broad Specificity
J. Immunol. 183:1207-1214. (2009) Kivonat -
József Kardos , Péter Závodszky , Yuji Goto , Gábor Náray-Szabó , László Gráf , Katalin Szilágyi , Orsolya Barabás , Anna Palló , Veronika Harmat , Péter Gál
Revisiting the mechanism of the autoactivation of the complement protease C1r in the C1 complex: Structure of the active catalytic region of C1r
Mol. Immunol. 45(6):1752-1760 (2008) Kivonat -
László Beinrohr , Veronika Harmat , József Dobó , Zsolt Lörincz , Péter Gál , Péter Závodszky
C1 inhibitor serpin domain structure reveals the likely mechanism of heparin potentiation and conformational disease
J. Biol. Chem. 282: 21100-21109. (2007) Kivonat -
Márton Megyeri , Veronika Harmat , Balázs Major , Ádám Végh , Júlia Balczer , Dávid Héja , Katalin Szilágyi , Dániel Datz , Gábor Pál , Péter Závodszky , Péter Gál , József Dobó
Quantitative characterization of the activation steps of Mannan-Binding Lectin (MBL)-Associated Serine Proteases (MASPs) points to the central role of MASP-1 in the initiation of the complement lectin pathway
J Biol Chem 288(13):8922-8934. | DOI: 10.1074/jbc.M112.446500 | PMID: 23386610 (2013) Kivonat -
Dávid Héja , Veronika Harmat , Krisztián Fodor , Matthias Wilmanns , József Dobó , Péter Závodszky , Péter Gál , Gábor Pál
Monospecific inhibitors show that both mannan-binding lectin-associated serine protease-1 (MASP-1) and -2 Are essential for lectin pathway activation and reveal structural plasticity of MASP-2
J. Biol. Chem. 287(24):20290-300. doi: 10.1074/jbc.M112.354332 | PMID: 22511776 (2012) Kivonat -
József Dobó , Veronika Harmat , Edina Sebestyén , László Beinrohr , Péter Závodszky , Péter Gál
Purification, crystallization and preliminary X-ray analysis of human mannose-binding lectin-associated serine protease-1 (MASP-1) catalytic region
Acta Crystallogr Sect F Struct Biol Cryst Commun. 64(Pt 9):781-784. | DOI: 10.1107/S174430910802294X | PMID: 18765903 (2008) Kivonat -
Péter Gál , Andrea Kocsis , Tünde Bián , Veronika Harmat , László Barna , Géza Ambrus , Barbara Végh , Robert B. Sim , Péter Závodszky , Gábor Náray-Szabó
A true autoactivating enzyme. Structural insight into mannose-binding lectin-associated serine protease-2 activations
J. Biol. Chem. 280(39):33435-33444. | PMID: 16040602 (2005) Kivonat -
Veronika Harmat , Péter Gál , József Kardos , Katalin Szilágyi , Géza Ambrus , Gábor Náray-Szabó , Péter Závodszky
The structure of MBL-associated serine protease-2 reveals that identical substrate specificities of C1s and MASP-2 are realized through different sets of enzyme-substrate interactions
J. Mol. Biol. 342(5):1533-1546. | PMID: 15364579 (2004) Kivonat